Awareness
DNA Day 2026
Decoding the Unseen: Molecular Fingerprints of the Environment
Molecular Microbiology and Human Diseases Project
Published on Thu, 04/23/2026 - 14:19

DNA Day
The global scientific community celebrates World DNA Day every year on 25th of April to commemorate both the anniversary of 1953 publication of DNA structure by Nobel prize-winning scientists James Watson and Francis Crick, together with Maurice Wilkins. This discovery was made possible by the pivotal X-ray diffraction data captured by Rosalind Franklin and the completion of the Human Genome Project (NHGRI, 2024) in 2003. These milestones in the field of science are not only framed as the foundation of molecular biology, but they also mark the beginning of a more significant change in our understanding of the environment. Now we understand that the overall well-being of our ecosystems and human society is determined by a huge, invisible library of genetic information on the molecular level (Kaplan, 2025; National Human Genome Research Institute, 2020).

Figure 1: James Dewey Watson; Francis Harry Compton Crick by Antony Barrington Brown, 1953. Credit: This Photo by Unknown Author is licensed under CC BY-NC
Figure 2: Rosalind Franklin observing through microscope, Credit: Science Source/SPL
Understanding the field of Molecular Microbiology
Over the last few decades, Microbiology has been revolutionized by the field of Molecular Biology, opening new research areas for scientists through the introduction of molecular techniques into microbiology research. Since the majority of microbes on earth remain unculturable using standard microbial culture techniques, there is a huge knowledge gap in how ecosystems function (NSR, 2021). DNA Sequencing, metagenomics and bioinformatics like advanced molecular techniques allow the identification of microbes directly from extracted DNA in different samples, including water, soil, air, blood etc. This capability has disclosed previously unknown levels of biological diversity. For instance, during the 2019 transboundary haze events in Sri Lanka, we found that the percentage of culturable bacterial species was high during the haze event compared to non-haze days and two human pathogenic bacteria were isolated and identified by using Sanger sequencing and bioinformatics tools (Thilakarathne et al., 2021). Moreover, our study conducted in the Mahapelessa hot spring in Sri Lanka utilized both culture-dependent and culture-independent metagenomic approaches to determine the microbial diversity in extreme environments. The V3-V4 amplicon metagenomic sequencing of DNA extracted from water samples allowed us to identify a total of 23 bacterial phyla representing 80 classes, 43 orders, 123 families, 205 genera and 83 species (Samarasinghe et al., 2021).
The evolution of Genomic Monitoring
Advanced sequencing methods allow us to monitor genomic variations more effectively than ever before. Bioinformatics serves as the mandatory bridge between raw, massive-scale sequences and actionable knowledge. Researchers can now identify specific genetic markers for antibiotic resistance, pathogenicity, or environmental adaptation with complex metagenomics datasets by utilizing advanced algorithms. Traditional culture based methods provide a foundational approach to capture microbes but often fail to capture the true breadth of microbial communities in our unique tropical ecosystems.
For over a century, the fundamental method of studying microbes was through culture dependent techniques, requiring scientists to grow microbes on a culture medium in a laboratory setting. Nevertheless, since over 99% of environmental microbes are unculturable, this created a huge knowledge gap in the field of microbiology. The incorporation of culture independent metagenomics was the first major shift in the microbiology research field, where DNA is extracted directly from environmental samples, bypassing the need for microbial culturing entirely. Once DNA is extracted from the samples, the Next-Generation Sequencing (NGS) phase is recommended. Metagenomic techniques allow researchers to identify entire microbial communities rather than focusing on the identification of a single microbe. The raw data generated by sequencers is essentially a massive, unresolved puzzle. The next most important stage in the identification of microbial community is Bioinformatics. The raw data collected after sequencing is assembled and compared against global databases using different computational tools and algorithms. This stage is where “raw data” becomes “identifiable sequences”. The final stage is real-time surveillance and policy integration where the flow of data leads directly to public health policy, acting as an early warning system.
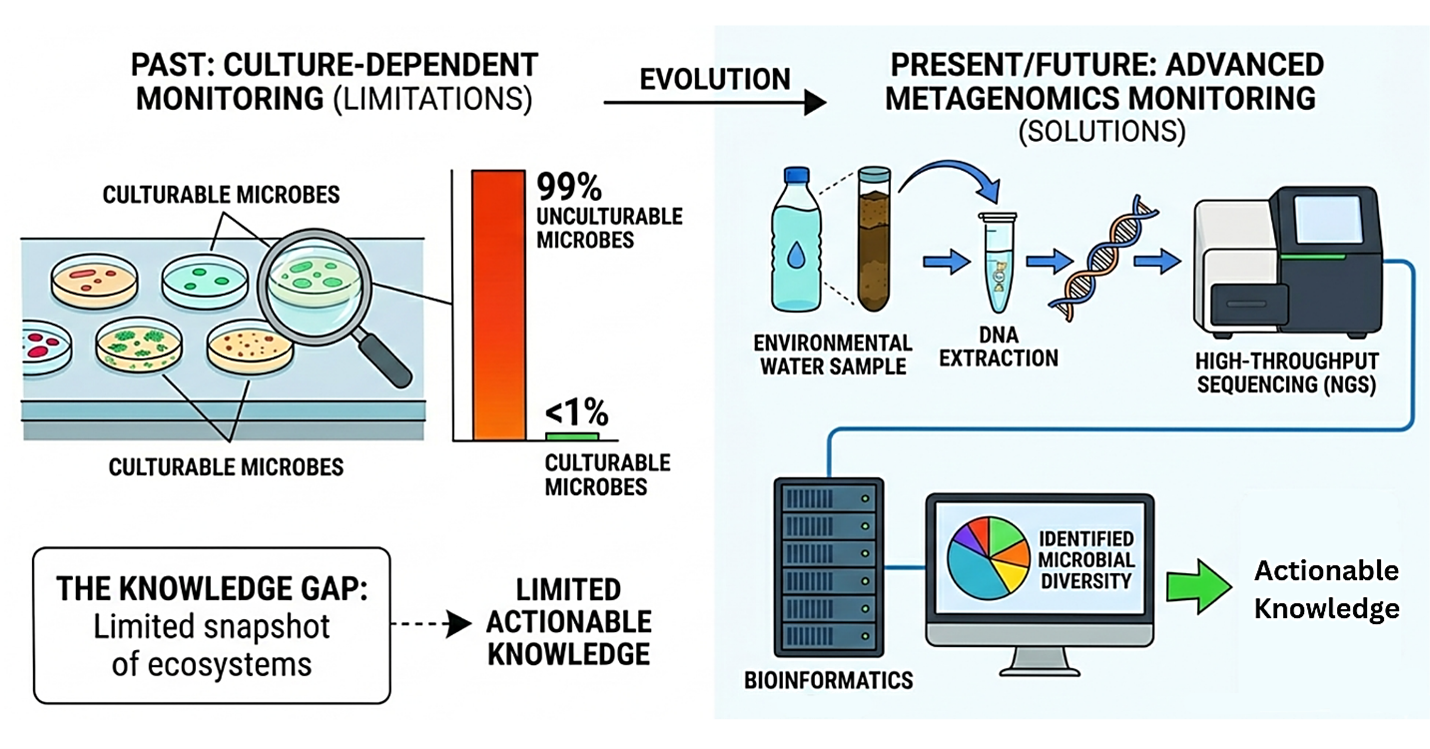
Figure 3: Evolution of Genomic Monitoring; Moving from traditional culture-based techniques to advanced metagenomics and bioinformatics for environmental and public health surveillance
Integrating knowledge on DNA for future resilience
Scientists are building a foundation for genomic foresight by using DNA technologies to decode the hidden information within our environments. This approach allows us to detect subtle and early warnings for shifts such as the accumulation of resistance genes in microbial genomes or the emergence of unknown or pathogenic microbial communities, before they become broad scale public health threats or lead to ecological collapse. In the 21st century, we are trying to bridge the gap between high resolution laboratory discovery and real world application. This means considering the environment as a dynamic and interconnected system where the flow of genetic information is constantly monitored. By establishing routine mapping of these "microbial fingerprints”, we can provide useful information that influences policy generation, guides resource allocation and develops an ability to adapt to environmental change.
As we celebrate DNA 2026, it is important to recognize the importance of fundamental science discoveries in providing practical, scalable solutions to real-world challenges. The genetic code embedded in our environment is both a narrative of our past and a warning for our future, interpreting this genetic information with precision is the most crucial task of modern science. When we standardize how to decode and read the genetic information of our planet, we acquire the capability to safeguard our communities against global environmental and health challenges, ensuring that the progress of science directly influences a more stable and prepared global society.
References:
- Kaplan, J. (2025, September 8). Francis Crick, Rosalind Franklin, James Watson, and Maurice Wilkins. Science History Institute. https://www.sciencehistory.org/education/scientific-biographies/francis-crick-rosalind-franklin-james-watson-and-maurice-wilkins/
- National Human Genome Research Institute. (2020). Homepage. National Human Genome Research Institute (NHGRI); Genome.gov. https://www.genome.gov/
- Samarasinghe, S. N., Wanigatunge, R. P., & Magana-Arachchi, D. N. (2021). Bacterial Diversity in a Sri Lankan Geothermal Spring Assessed by Culture-Dependent and Culture-Independent Approaches. Current Microbiology, 78(9), 3439–3452. https://doi.org/10.1007/s00284-021-02608-4
- Thilakarathne, S. M. N. K., Ekanayake, A., Madamarandawala, P. S., Weerarathne, W. B. C. P., Thotawatthage, C. A., & Magana-Arachchi, D. N. (2021). Impact of haze events on airborne bacterial consortia–a case study. SN Applied Sciences, 3(1). https://doi.org/10.1007/s42452-020-04022-0